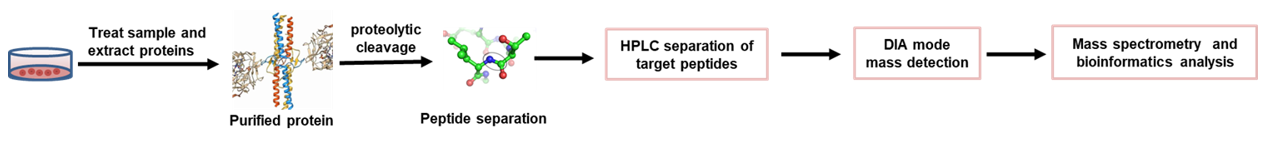
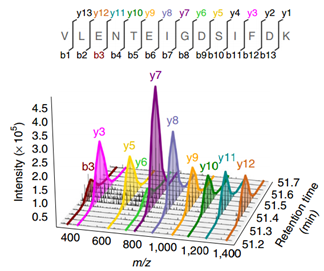
A data-independent acquisition-based global phosphoproteomics system enables deep profiling | Nature Communications

Thulasi Its Pro Dia 200gm Protein Shake Price in India - Buy Thulasi Its Pro Dia 200gm Protein Shake online at Flipkart.com

Going library-free for protein identification using Zeno SWATH data-independent analysis (DIA) and in silico-generated spectral libraries

Direct data-independent acquisition (direct DIA) enables substantially improved label-free quantitative proteomics in Arabidopsis | bioRxiv
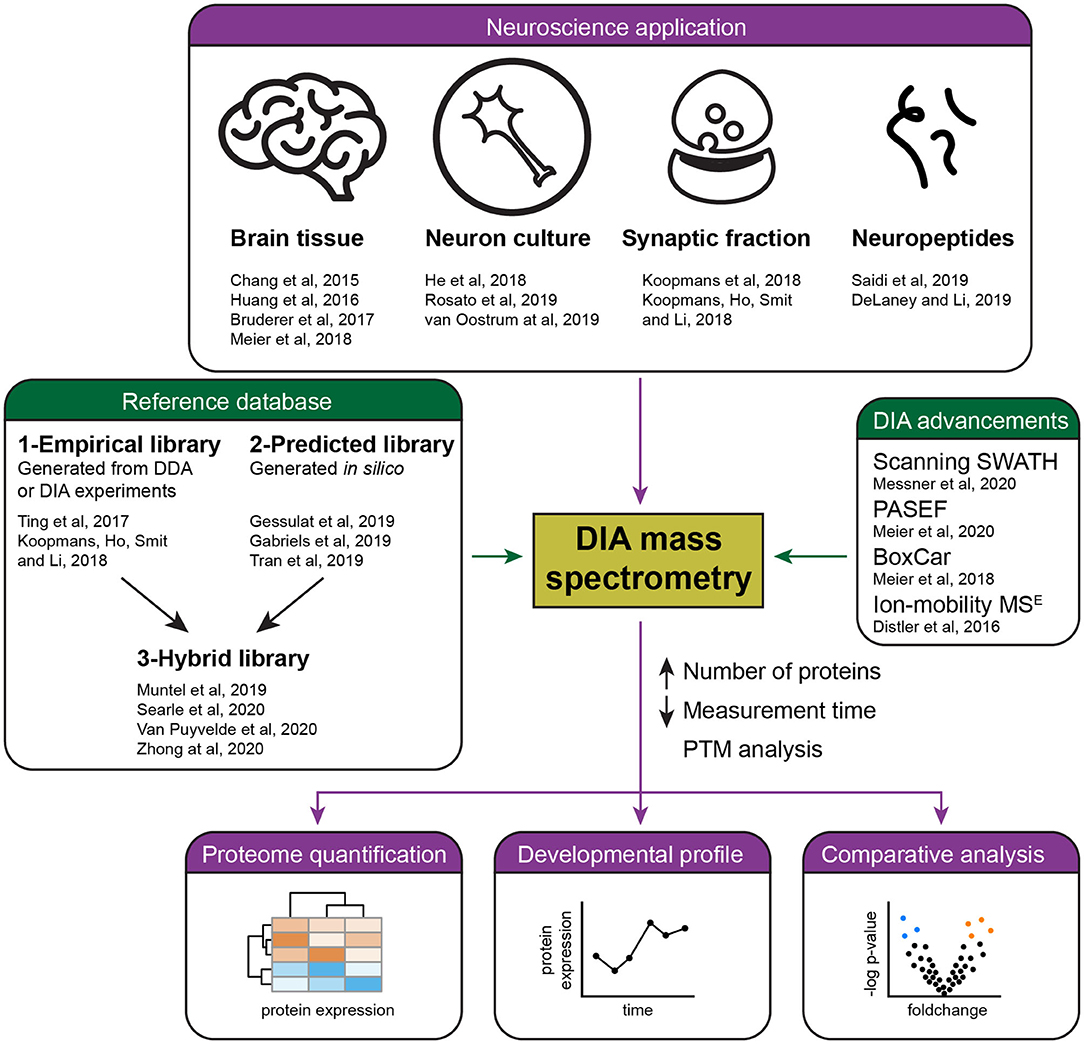
Frontiers | Recent Developments in Data Independent Acquisition (DIA) Mass Spectrometry: Application of Quantitative Analysis of the Brain Proteome
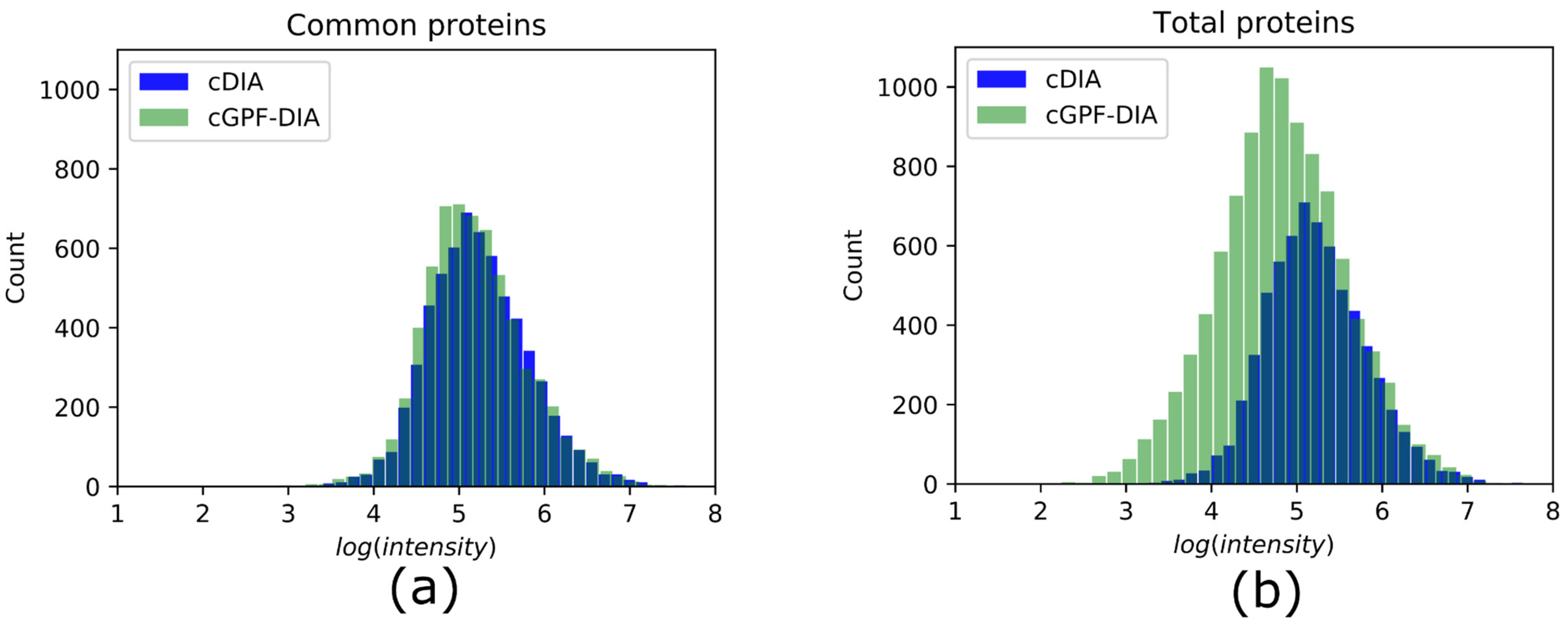
Life | Free Full-Text | Narrow Precursor Mass Range for DIA–MS Enhances Protein Identification and Quantification in Arabidopsis

Direct data-independent acquisition (direct DIA) enables substantially improved label-free quantitative proteomics in Arabidopsis | bioRxiv

Improved SILAC Quantification with Data-Independent Acquisition to Investigate Bortezomib-Induced Protein Degradation | Journal of Proteome Research

Parallel Factor Analysis Enables Quantification and Identification of Highly Convolved Data-Independent-Acquired Protein Spectra - ScienceDirect

Protein Contaminants Matter: Building Universal Protein Contaminant Libraries for DDA and DIA Proteomics | Journal of Proteome Research

Dia-Interacting Protein Modulates Formin-Mediated Actin Assembly at the Cell Cortex: Current Biology
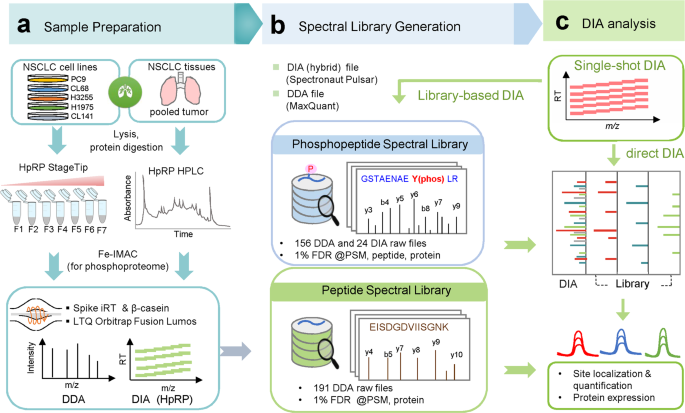
A data-independent acquisition-based global phosphoproteomics system enables deep profiling | Nature Communications

Dia dependent cellular structures and functions. Members of Diaphanous... | Download Scientific Diagram
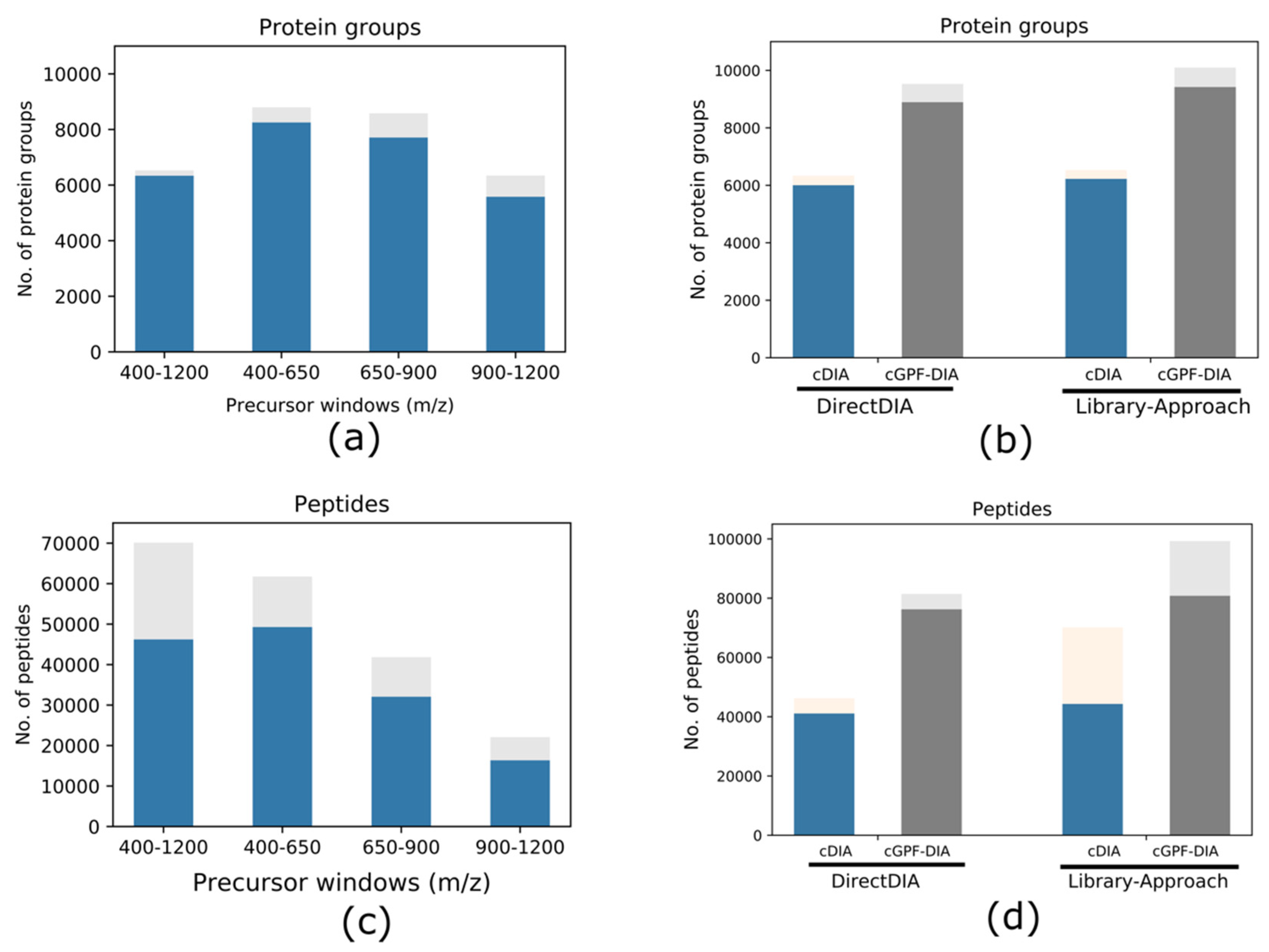
Life | Free Full-Text | Narrow Precursor Mass Range for DIA–MS Enhances Protein Identification and Quantification in Arabidopsis